Computational Notebook
18 November 2020
A computational notebook is an environment for writing a prose document that allows the author to embed code which can be easily executed with the results also incorporated into the document. It's a platform particularly well-suited for data science work. Such environments include Jupyter Notebook, R Markdown, Mathematica, and Emacs's org-mode.
When I'm exploring some data, it's useful to keep my notes close together with the code that performs the exploration. I like to try some code, look at the results, and note down any observations I have from that execution. A computational notebook allows me to combine these together easily in a single document.
Here's an example of this, looking at some analysis of my google analytics data for martinfowler.com. I'm doing this in R Studio, which uses the R Markdown format.
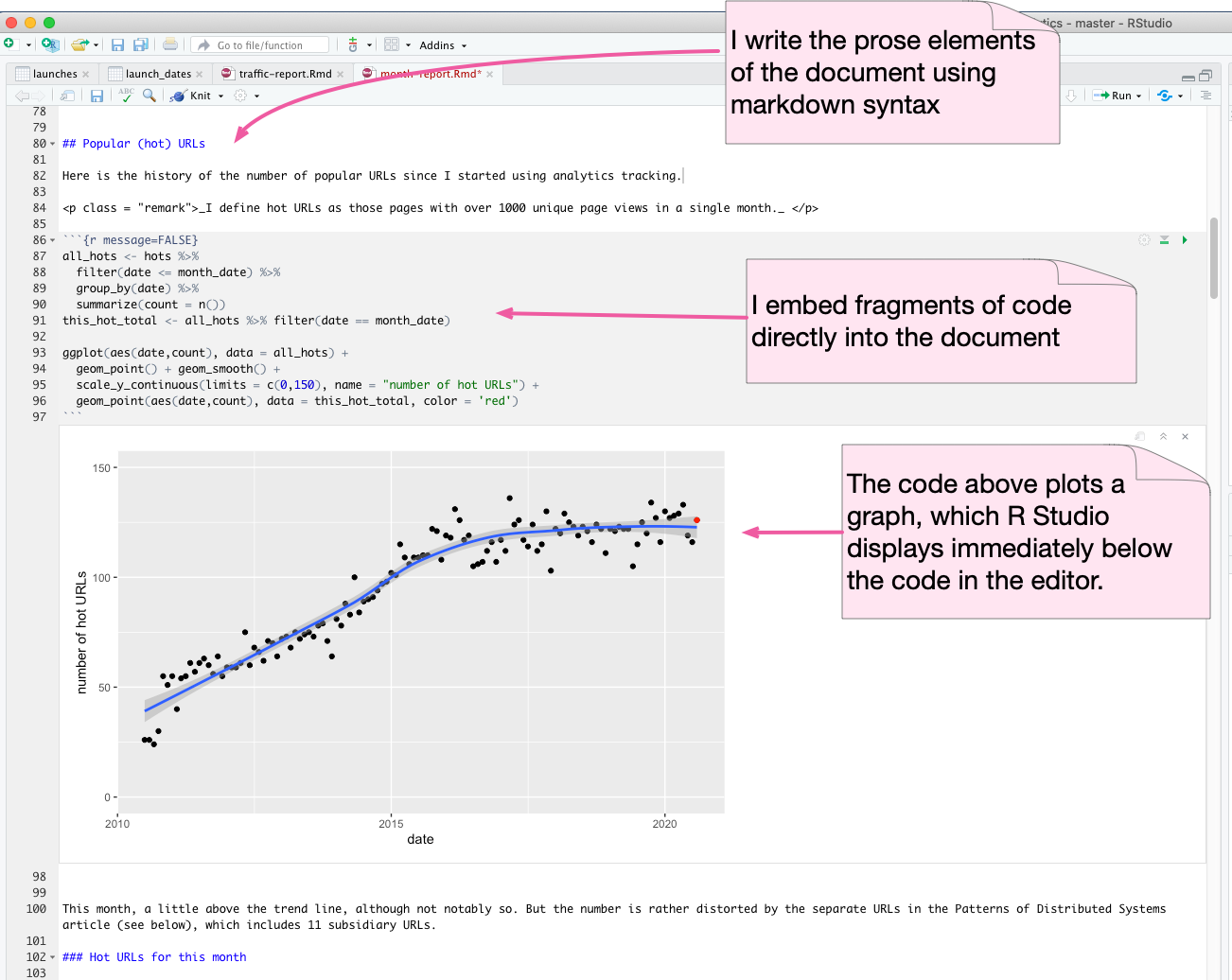
The example out here is a graph, as notebooks are well suited for plotting various charts. But it's just as useful to embed various data manipulations in the code and display the data in the document as a table.
I first encountered a computational notebook in the late 1980's with Mathematica. I remember wishing I'd had access to such a tool during my university degree, but didn't use a computational notebook again until recent years, with the rise of their use in data science circles. The notebook software I hear most about is Jupyter Notebook, which is popular in the Python community, but as I do my data munging with R I tend to use R Markdown, usually within R Studio. I also use a rather more niche notebook, org-mode, which is part of Emacs.
The code embedded in Mathematica is its own programming language, designed for expressing mathematics. Although Jupyter began in the Python world, it supports a wide range of programming languages, as does R Markdown. Mathematica is a commercial tool, but Jupyter and R Markdown are open source. Jupyter stores its files in JSON, R Markdown uses markdown files with some special markup for the code blocks. Using a text format for the documents allows them to be stored in regular version control tools, and using a markup language makes diffing easier. Using a markup language allows the possibility of editing the documents in other editors, but they need to have a suitable environment for executing the code blocks.
Computational notebooks are useful when exploring a problem, such as trying various forms of analysis on a dataset. The document acts as a record of what's been tried and all the observations the researcher makes as they try things. By keeping the code and results together the writer can see exactly what they did and what results that generated. This coupling of code and results is a form of IllustrativeProgramming, making the environment appealing to lay programmers. One thing to be wary of, however, is if any external environmental factors change the result - such as the contents of a database. If the dataset isn't too large it can be exported and kept in the version control system, but often its size is prohibitive.
Notebooks are also useful for preparing reports, usually by generating a document in PDF, HTML, or other formats. If I want to report to an author on the traffic for their article, I take the last such report, change the subject URL, rerun all the code, and tweak any prose commentary I think is appropriate. If I were sufficiently motivated I could auto-generate such reports every few months. I like that such reports can easily include the code used to generate the results, so readers can accurately understand the logic behind the figures they see.
Notebooks shouldn't be used, however, as a component of a production system. The notebook structure - with its casual mix of IO, calculation, and UI - is there to encourage interactivity, but works against the modularity needed for code that is used as part of a broader code base. It's best to think of notebooks as a way of exploring logic, once you've found a path, that logic should be replicated into a library designed for production use.

